Improve sensitivity and specificity of eDNA metabarcoding tools for Invasive Marine Species Detection
eDGES v1 – Project 4
In this project we have enhanced and validated the use of metabarcoding of environmental DNA as a sampling tool for detecting Invasive Marine Species (IMS). This research project fits into a larger state-wide array surveillance program (SWASP) that was developed by the Department of Primary Industries and Regional Development (DPIRD) in consultation with the port authorities, shipping industry and the oil and gas sector in Western Australia.
Environmental DNA has advantages over conventional sampling in that it can detect cryptic, rare and early life stage species in water and biofouling that a person may not readily observe. However, eDNA is a relatively new tool that needs further validation, particularly in the IMS area. It is recognised that the confidence in IMS detections is only as good as the ability of the tests and the genetic reference sequences that they are based on.
This project has contributed to the global reference databases through barcoding target IMS and their native congeners. We also developed and validated new metabarcoding assays that are able to assign species identity to genetic sequences with a high level of confidence, allowing the detection of target IMS from a range of biological substrates.
Overview of outputs and outcomes achieved to date
The first stage of Project 4 aimed to fill gaps in the molecular data available for priority IMS and their native taxa, in some cases there are limited, or no sequence data from taxonomically confirmed specimens.
- Finalised GenBank submission of 31 full mitochondrial genome sequences, 7 partial mitochondrial genomes and five chloroplast genome sequences.
- Compiled a custom 16S rRNA reference database of native Western Australian congeners to improve taxonomic assignment from eDNA sampling. This was achieved by sequencing the 16S rRNA gene region of samples acquired from DPIRD’s collections and specimens collected during field trips to Dampier (West side of the Burrup Peninsula, Oct 21), Onslow (Aug 22), Dampier (East side of the Burrup Peninsula and islands in the Dampier Archipelago, Oct 22) and Port Hedland (Jan 23). Available sequence data obtained from GenBank, DPIRD and WAM databases was also included in this reference database.
Wells FE, Lukehurst SS, Fullwood LAF and Harvey ES (2024) Distribution of intertidal rock oysters in the Pilbara, Western Australia. Management of Biological Invasions 15: 131-143.
- A previously unknown species of Talonostreahas been found in Hooley Creek, the Dampier Archipelago and at Balla Balla. The species is small, about 3cm in diameter and occurred in three intertidal habitats: on sand in a creek, attached to the underside of mangroves and on a steel structure. The Pilbara DNA sequences match a new species of Talanostrea recorded by Dr Carmel McDougall of Griffith University between Brisbane and Cairns. Dr Carmel McDougall is describing the new species with Dr Graham Oliver of the National Museum of Wales. The Pilbara data will be included in this species description.

Figure 1: Newspaper article discussing the newly identified Talonostrea sp. This article was also featured in the online West Australian and the North West Telegraph, May 2024.
The second stage of the project was to develop novel metabarcoding assays, to enable discrimination between IMS and closely related native species within phyla Arthropoda and Mollusca.
- Two assays, targeting the 16SrRNA region, were designed to target both Arthropoda and Mollusca. Assays were initially tested onsingle sourcetissue gDNA extracts from IMS of interest and native congeners. This single source gDNA was also combined to create mock communities, which were used as positive controls for the metabarcoding workflow. The IMS of interest were successfully detected in these mock communities using the relevant assay.
- We tested the assays using eDNA extracted from biofouling from settlement arrays deployed at Dampier port. In the environmental samples, the Arthropoda assay detected 12 genera from Malacostraca, three genera fromHexanaupliaand one from Insecta. The Mollusca assay detected 15 genera from Bivalvia and 10 genera from Gastropoda. Even though the Mollusca assay was able to detect 25 genera, 97.4% of sequence reads were unassigned, this indicates the importance of reference sequences and updated databases.
- The Arthropoda assay was used to screen eDNA extracted from water and plankton tow samples collected during DPIRD’s routine surveillance forCharybdis japonicain the Swan River, between 2019 and 2022. The assay had detections of Arenallianassa arenosa, Macrobrachium intermedium, Panulirus cygnus, Penaeus latisulcatus, Portunus armatus, Thalamita sima, Balanus trigonus, Armatobalanus sp., Tetraclitella purpurascens. All common species found in the area, with a greater number of sequence reads found in the plankton tow samples.
- To determine the sensitivity of the assays, two synthetic oligonucleotides were designed to be used as positive controls. These positive controls were based on IMS 16S gene regions with a few changes to allow discrimination from actual Balanus glandula and Perna viridis DNA. The synthetic positives were spiked into DNA extracts from three different environmental substrates (settlement array, water filter, plankton tow) at three different concentrations. The Mollusca assay was more sensitive than the Arthropoda assay with 4 copies/mL being detected in spiked samples when assay was run as either a single- or multi-plex. The lowest concentration of 4.7 copies/mL was detected in 66% of spiked samples when the Arthropda assay was performed in a multiplex, otherwise 47 copies/mL was the limit of detection when the assay was performed alone.
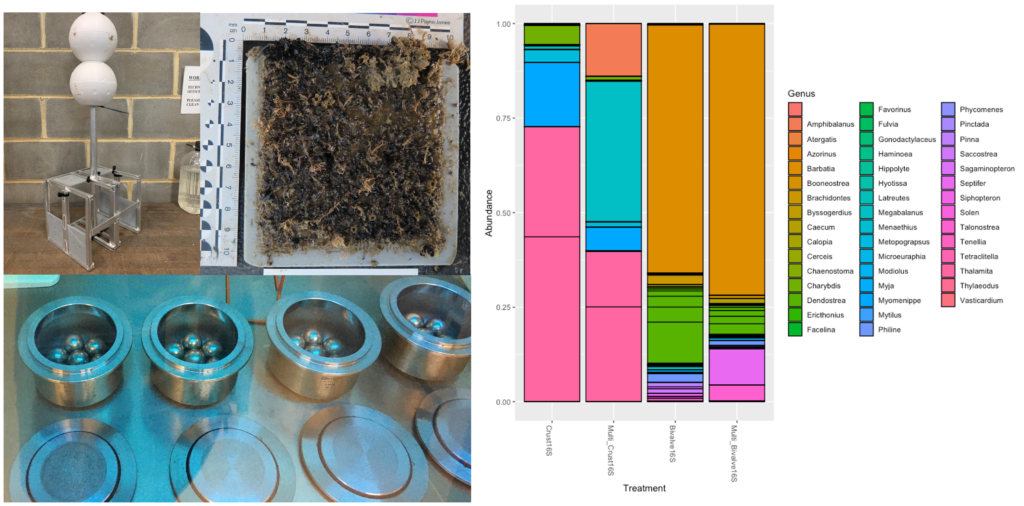
Figure 2: DNA extracted from biofoul that settled onto plates deployed at Dampier port was screened with the new assays in singleplex and multiplex.
Additional projects:
- As part of eDGEs1 we collaborated with Dr Tan Koh Siang from the National University of Singapore on species of the Brachidontes variabilis complex in the Indo-West Pacific and published the results. DNA extractions and sequencing of five gene regions for specimens from three sites in WA (Dampier, Woodman Point and Albany) have been completed and submitted to GenBank.
Tan SHM, Wells FE, Lukehurst SS, Strong E, Sanpanich K, Duangdee T, Ambarwati R and Tan KS (2024) Unravelling the Brachidontes variabilis species complex (Bivalvia: Mytilidae). Journal of Molluscan Studies 90: eyae037, 23 pages.
- There have been extensive vessel movements between Singapore and Thailand and the WA Pilbara region by vessels that are high risk for introducing invasive marine species. In particular, the invasive oyster Magallana gigas has been reported in the scientific literature from both countries. In February 2023 Dr Wells and Ms Lukehurst went to Singapore to work with Dr Tan Koh Siang and Tan Siong Kiat of the National University of Singapore to clarify the identity of Singapore oysters and the IMS risk they pose to Australia. Two genera were surveyed: Magallana and Saccostrea. The field sampling included 27 sites. Ninety tissue samples of oysters and mussels were brought back to Curtin for processing. Shells and remaining tissue of all specimens were retained by Lee Kong Chian Natural History Museum and have been vouchered. DNA extractions, PCRs, sanger sequencing and sequence editing has been completed for all specimens.
- A manuscript reporting on species of the oyster genus Magallana in Singapore and their biosecurity risk to Australia has been published. M. gigas does not occur in Singapore but two related species, M. belcheri and M. bilineata do occur there.
Tan KS, Tan SK, Lukehurst S and Wells FE (2024) Absence of the Pacific oyster Magallana gigas (Thunberg, 1793) in Singapore. Raffles Bulletin of Zoology 72: 162-183.
- As with Singapore, we undertook a survey of intertidal oysters in the Gulf of Thailand in collaboration with Dr Teerapong Duangdee (Kasetsart University) and Dr Kittithorn Sanpanich (retired from Burapha University). We published a paper on the genera Magallana and Saccostrea in the northern Gulf of Thailand. The survey enabled us to also investigate and publish on the current status of the invasive charru mussel Mytella strigata five years after it was first identified in the Gulf of Thailand. The charru mussel poses a significant IMS threat to Australian waters.
Wells FE, Duangdee T, Sanpanich K & Lukehurst SS. (2024) Status of the invasive charru mussel Mytella strigata (Hanley, 1843) in the upper Gulf of Thailand five years after it was first surveyed. BioInvasions Records 13: 69-82.
Duangdee T, Sanpanich K, Lukehurst SS & Wells FE (2025) Oysters in the genera Magallana and Saccostrea in the upper Gulf of Thailand based on 16SrNA DNA sequences. Raffles Bulletin of Zoology 73: 12-33.
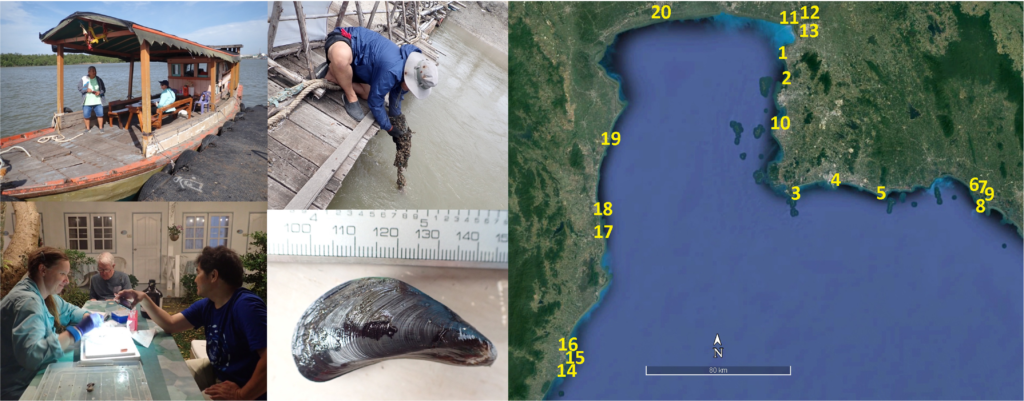

Figure 3: Mytella strigata survey in the upper Gulf of Thailand, August 2023. Yellow circles represent sampling sites, with closed circles indicating the sites where the species was present at the time of the survey.
- Research into possible assays that can be used to distinguish IMS of macroalgae from closely related native species. Macroalgae specimens have been acquired from DPIRD and DBCA Herbarium collections. DNA has been extracted and the 23S chloroplast gene region sequenced to create an algae reference database for this region. For the genus Sargassum there was not enough variation between species, therefore other gene regions were considered. Primers for the space between the rbcL and rbcS chloroplast gene regions have been designed and are awaiting testing.
