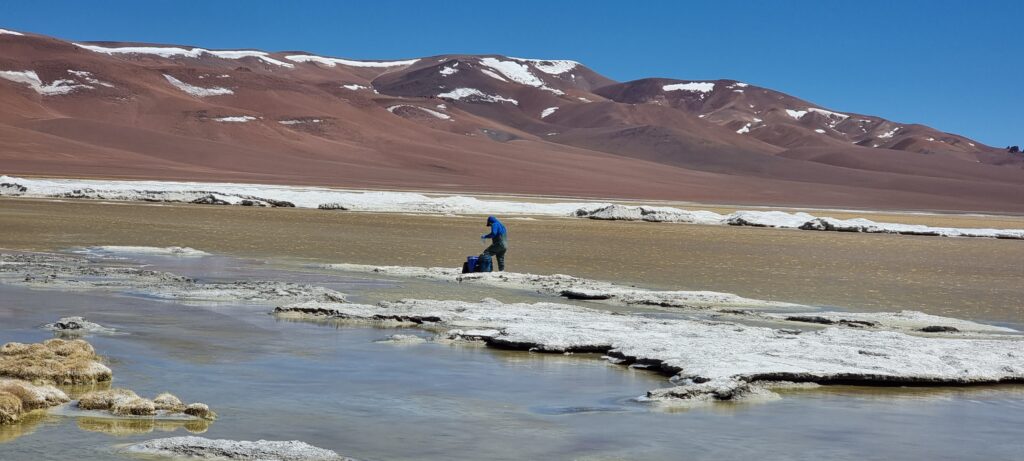
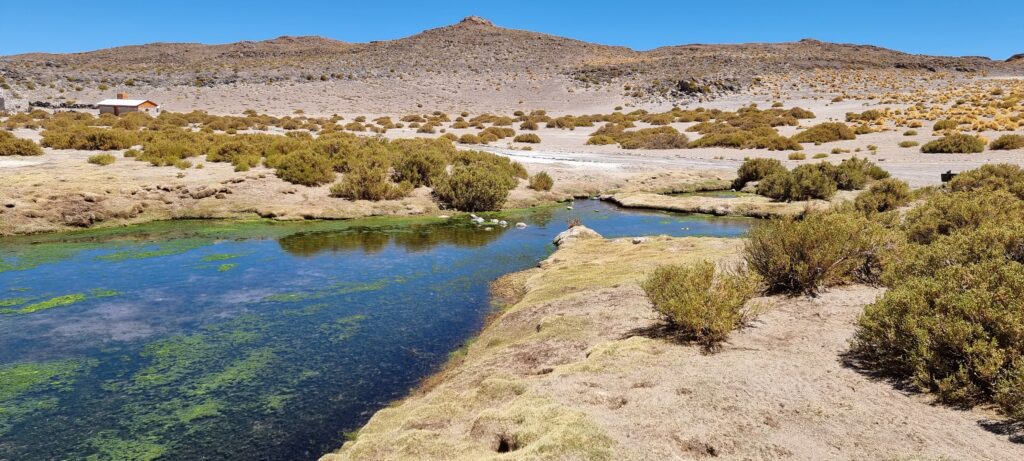
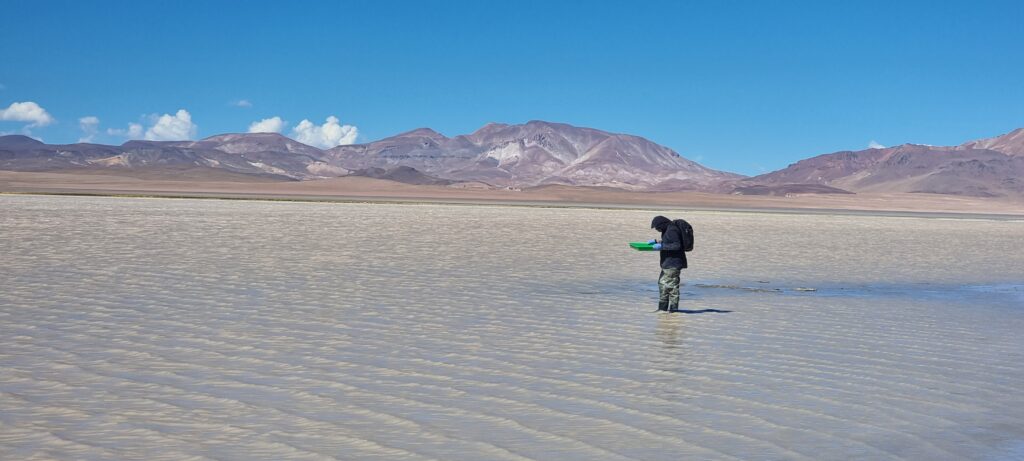
Functional ecology of Chilean precordillera lakes: towards a Wetland Health Index for birds
eDGES v1- Project 3
In this project we used eDNA technology (and isotope analysis) to monitor biodiversity and characterize ecosystem functioning of hypersaline lakes within the Chilean precordillera. These environments provide habitats and resources that are key to the survival of several species of high conservation concern, including the Chilean James’s and Andean flamingos. However, the current knowledge of the ecological dynamics that sustain these hypersaline wetlands is poor. This study incorporated eDNA monitoring for a wide range of prokaryotic and eukaryotic taxa in water and sediments, combined with biogeochemical investigations on the aquatic ecosystems targeted . This approach establishes a solid knowledge baseline in understanding the ecosystem functioning that is shaping key water bird habitats such as the high-altitude hyper-saline environments of the Chilean precordillera. By consistently widening current knowledge on diversity and functions in hypersaline lakes, the analytical toolkit developed through this project has the real potential to improve the coexistence of economic (i.e., mining) and non-economic (i.e., ecosystem) services into the future.
A literature review on the ecology and preservation of hypersaline wetlands was published in the prestigious Journal Biological reviews in 2021. (https://doi.org/10.1111/brv.12780 ). The work developed through the review has also led to international collaborations with experts in Russia, China and Europe.

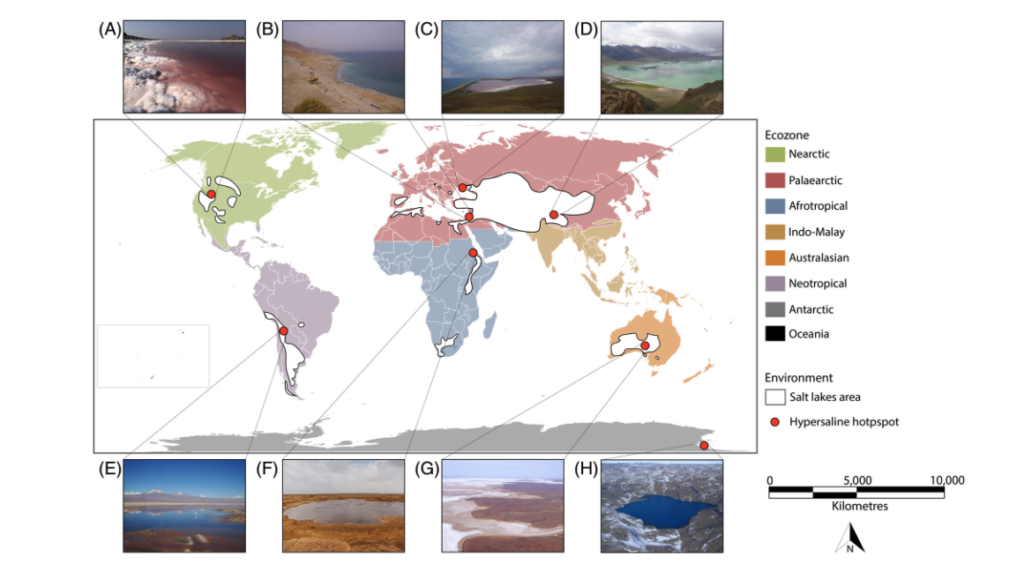
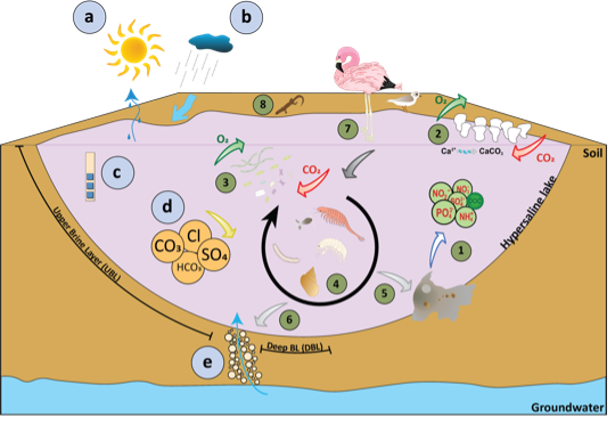
Figure 1: Front page and two graphs on the distribution and ecology of hypersaline lakes from the published manuscript.
Satellite investigations at Rottnest Island
A pilot study targeting the hypersaline lakes at Rottnest Island was initiated in 2020 with the goal of testing the state-of-the-art molecular techniques that were planned to be implemented in the Chilean precordillera. Several fieldtrips were carried out to assess the sampling techniques and finally collect samples from five hypersaline lakes on the island. The research associated with this satellite investigation has led to the publication of two manuscripts in Water (https://doi.org/10.3390/w13141899 ) and Journal of Oceanology and Limnology (https://doi.org/10.1007/s00343-022-2151-9) , respectively:
Metabarcoding under Brine: Microbial Ecology of Five Hypersaline Lakes at Rottnest Island (WA, Australia)
In this study we tested the use of eDNA techniques to assess the microbial communities of five hypersaline lakes at Rottnest Island. Our findings confirm the high effectiveness of DNA metabarcoding analysis in elucidating differential microbial patterns (diversity and functions) under different environmental conditions.
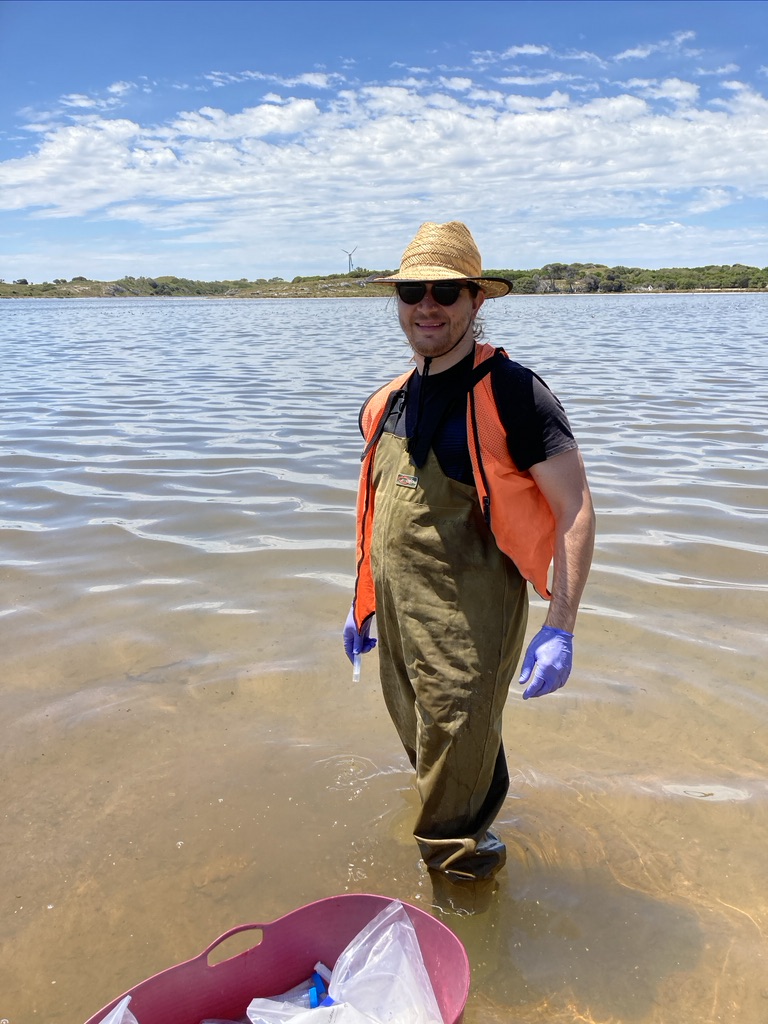

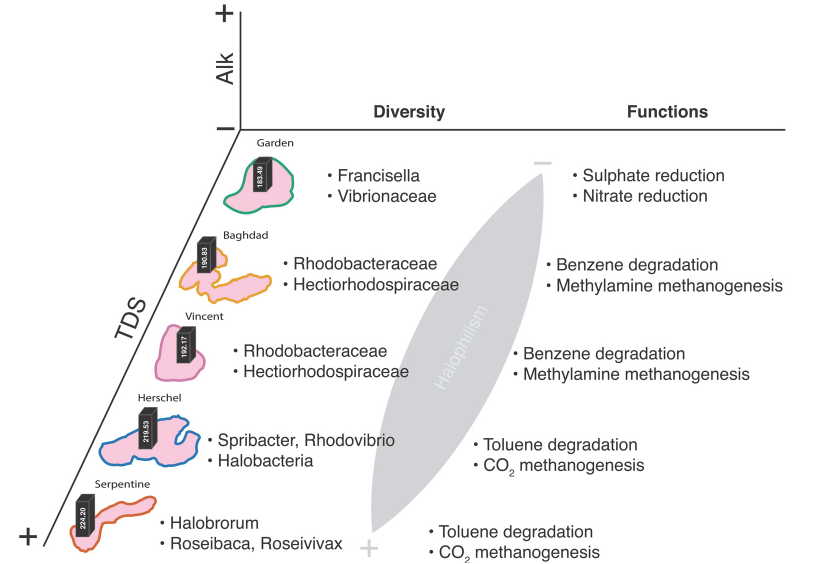
Figure 2: Photo from the fieldwork activities and summary graph from the published manuscript (https://doi.org/10.3390/w13141899 ) showing the diversity any functional patterns unveiled through eDNA techniques.
When nets meet environmental DNA metabarcoding: integrative approach to unveil invertebrate community patterns of hypersaline lakes
Here, data on macroinvertebrate diversity was compiled to compare results from macroinvertebrate conventional sampling techniques (nets) and eDNA approaches. Outcomes of this study suggest that eDNA tests provide a very valuable complementary tool able to detect taxa that are otherwise omitted via conventional approaches.
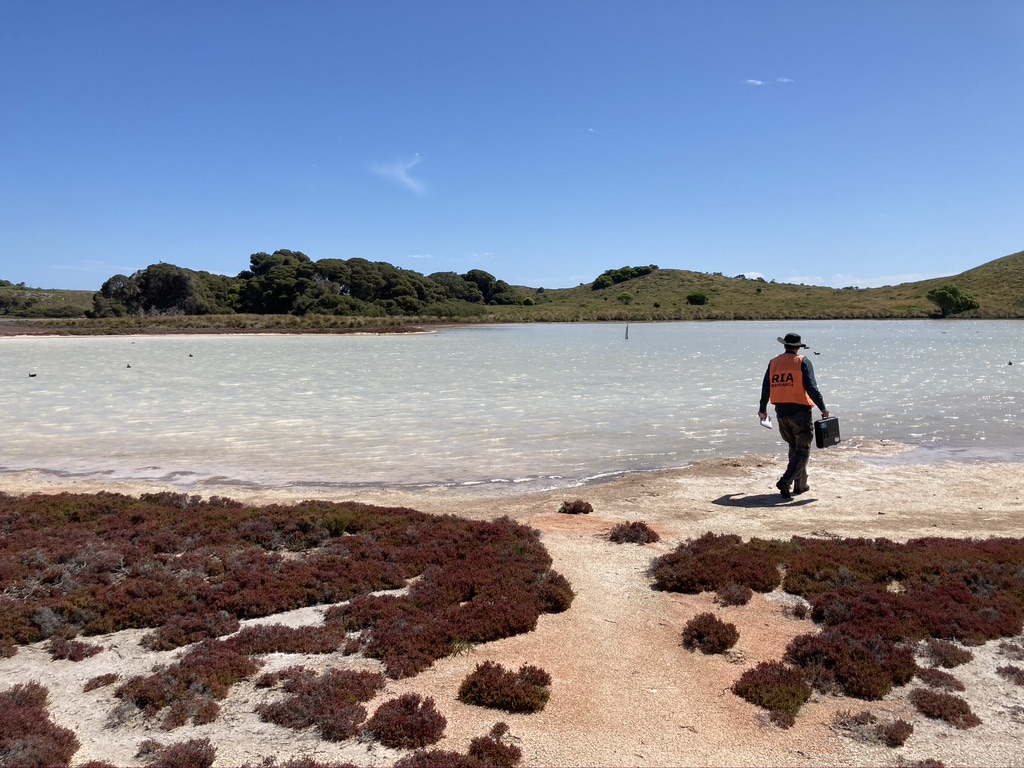

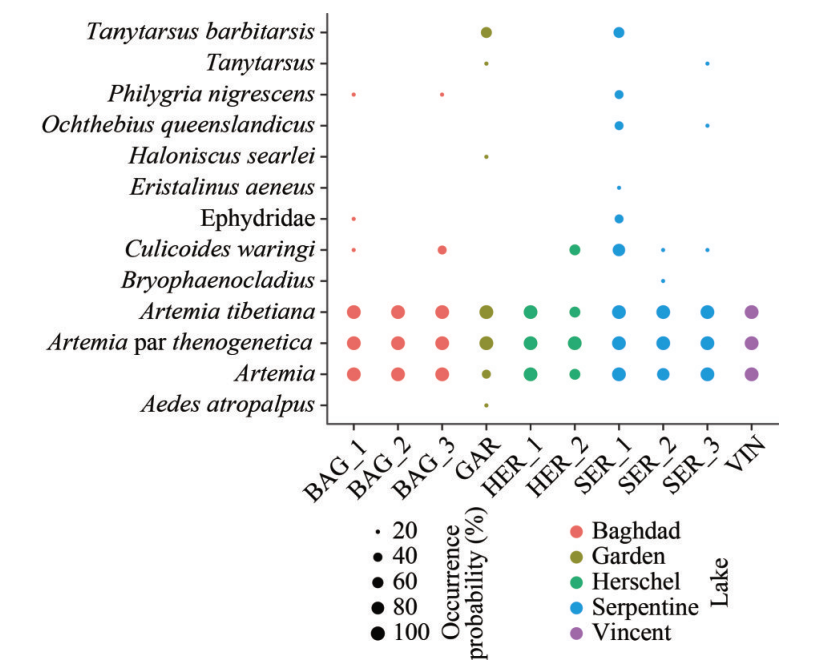
Figure 3: Photo from the fieldwork activities and graph from the published manuscript (https://doi.org/10.1007/s00343-022-2151-9) displaying the occurrence probability of macroinvertebrate taxa detected with DNA metabarcoding.
Chilean precordillera study
This was the main component of the project and involved the design, collection and analysis of samples from three hypersaline lakes located at different altitude in the Chilean precordillera: Puilar lagoon (Salar de Atacama), Salar de Tara and Salar de Pujsa. Fieldwork was carried out in October 2021 and both sediment and water samples were collected from each Salar. The vast dataset generated through DNA metabarcoding approaches has been published in Frontiers in Ecology and evolution:
Environmental DNA as a tool to unveil metazoan diversity of Chilean Precordillera hypersaline lakes
In this work (short communication) we investigated the use of eDNA analysis as a tool to unveil metazoan hypersaline diversity. Our results indicated that our multi-assay approach can unveil a vast group of endemic taxa (from microscopic to wading birds) across the three Salars targeted. Some examples of charismatic and hypersaline resident taxa detected through eDNA include flamingos (Phoenicopterus), gastropods (Heleobia atacamensis), amphipods (Hyalella), copepods (Diaptomidae) and ostracods (Heterocypris salina).
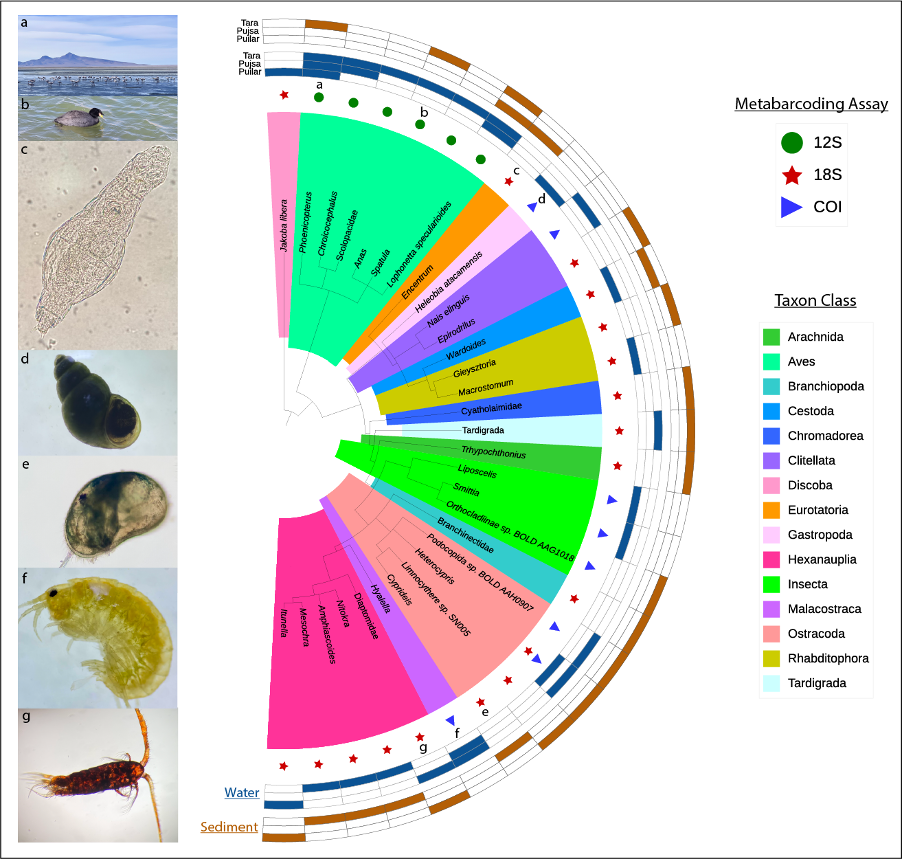
Figure 4: Graph from the manuscript (https://doi.org/10.3389/fevo.2025.1504666) showing the array of species found in water and sediment samples across the three Salars via DNA metabarcoding techniques.
Diversity of resident prokaryotic and eukaryotic microorganisms in three hypersaline lakes via DNA metabarcoding
In this work we provide an in-depth characterisation of the microorganismal diversity patterns of the three Salars investigated. A multi-assay approach (Bact16S – microbes, COI – fungi and protists, 18S – fungi protists, and rbcl – algae). Our results indicate that the Salars are shaped by high intra- and inter-lake microorganismal diversity patterns, highlighting the uniqueness of these ecosystems in terms of functional adaptations to high altitude and UV ranges.
Isotope work – The analysis of food web interactions and diet preferences has been led by Prof Chris Harrod and his team at the Universidad de Antofagasta Stable Isotope Facility (UASIF). Transfer of funds for analytical purposes was included in the fieldwork allowance (initially planned to cover two campaigns, but eventually allocated for just one sampling event in October 2021). Results are expected to be generated over 2025 and 2026, and data will be analysed and presented in Gonzalo’s Thesis and through publications in collaboration with the eDGES team of researchers.
Snapshots from the project



