Software
Tools for Environmental DNA Analysis and Data Interpretation
Seqma
eDNA metabarcoding produces results with a quasi-quantitative relationship between molecular abundance and the biomass of organisms represented by DNA barcodes. The relationship is imperfect for two types of reasons: stochastic factors and consistent biases.
Stochastic factors affecting the relationship between counts of DNA sequence types and the organisms they represent largely stem from eDNA for macro-organisms being sampled from near the organisms, rather than sampling them directly as is done for microbes. Consistent biases include differences in DNA:biomass ratios among organisms, variable rates of eDNA shedding, and systematic failure of PCR to amplify specific barcodes. Stochastic factors can be addressed by increasing sampling replication, whereas consistent factors must be addressed by controlled experiments.
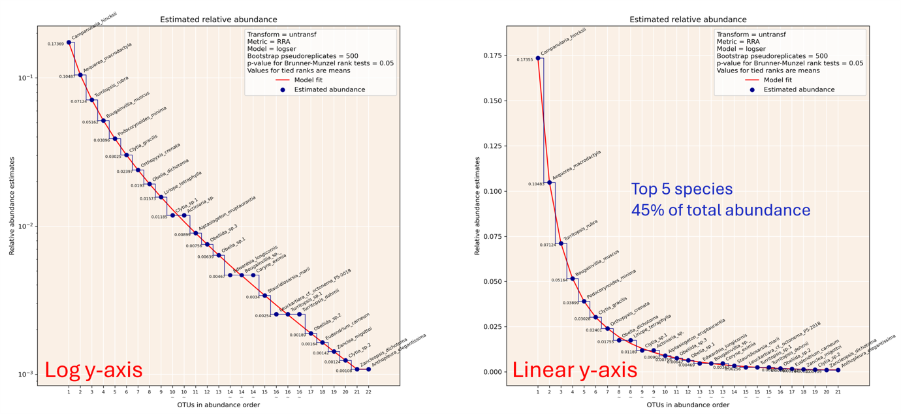
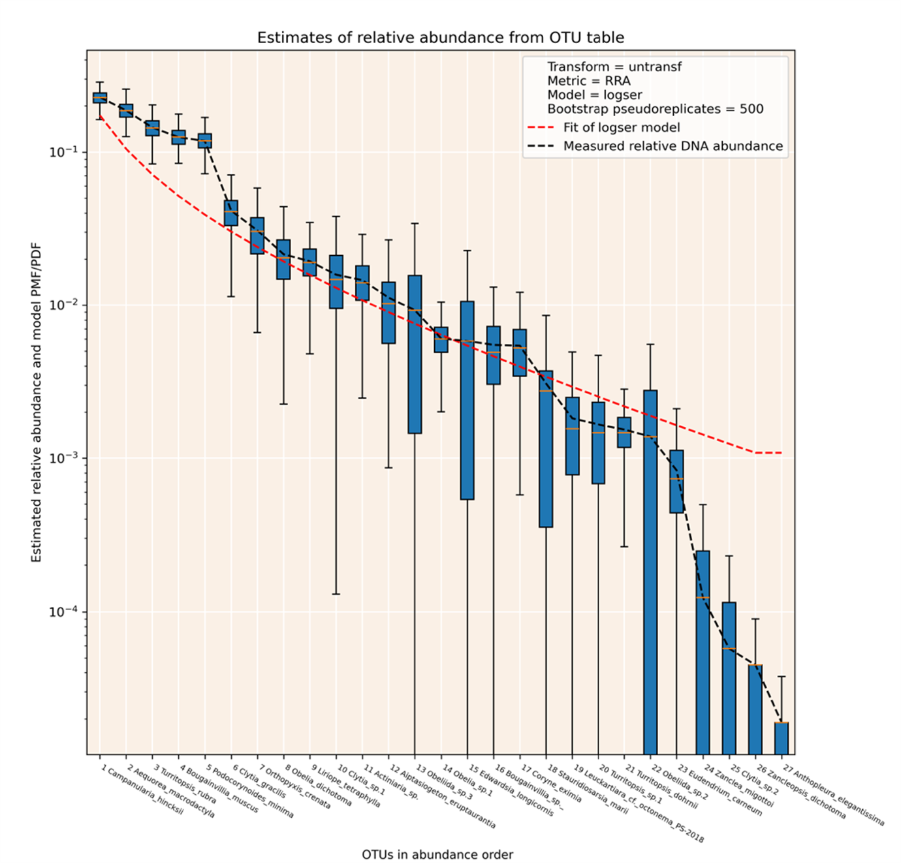
As part of project 8, software package for estimating the parameters of species abundance distribution (SAD) models from eDNA metabarcoding data has been written. This programme takes OTU table data as its input, which is a standard format for eDNA metabarcoding. Every OTU in the dataset is ranked by DNA read abundance, and reads are aggregated by one of a range of metrics. SAD curves are fitted to the rank abundance:pseudoabundance values, and the best fitting SAD curve is taken as the model. Species’ abundances can then be calculated from the SAD model. The Python package ‘SQEMA’ has been published on the python package index: https://pypi.org/project/sqema/

eDNA mapping tool
An interactive eDNA mapping tool is currently in development through a collaboration with eDNA Frontiers. This tool will allow users to explore, visualise, and interrogate environmental DNA data collected across a range of sites and taxa. Designed with accessibility and usability in mind, it enables filtering by location, species, and metadata fields, and supports on-the-fly data summaries. Users will also be able to download selected datasets for further analysis, providing what is hopefully a valuable resource for researchers, land managers, and citizen scientists interested in biodiversity patterns and ecosystem health.
